Background
Leukemia is a cancer that affects blood-forming tissue and therefore can be diagnosed based on the morphology and counts of certain white blood cells. Manually counting white blood cell subtypes can be an extremely tedious, time consuming, and inaccurate process for pathologists (Alomari et al., 2014). Additionally, being able to rapidly and accurately identify white blood cells can aid in medical research and the diagnosis of other diseases. This motivated us to develop deep learning models for white blood cell identification with the aim of diagnosing acute myeloid leukemia (AML).
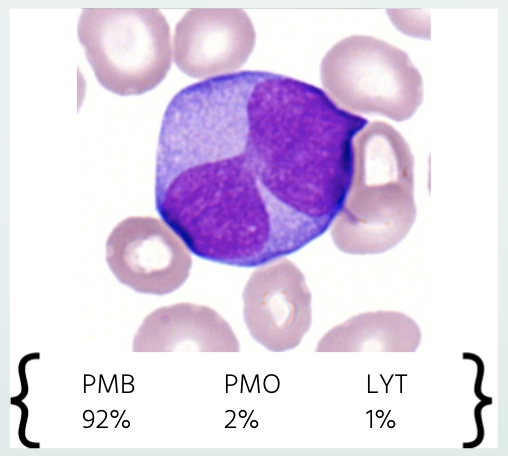
One issue with current machine learning research for leukemia diagnosis has been the lack of prospective applications of algorithms in real-world scenarios (Salah et al., 2019). Researchers predominantly use small and homogenous samples to train models that provide point predictions without explanation. These are some of the issues we had in mind while designing our white blood cell identifier. We explore a methodology from Angelopoulos et al., to quantify the uncertainty of our models’ predictions and provide accurate confidence intervals so that pathologists can understand the probability of certain cells being present. This feature can be useful in real world scenarios in multiple ways. For example, given a cell to identify, a pathologist can know with 95% confidence that the cell presented is not a myeloblast, which could be a cell indicative of AML if found in high numbers (DiNardo et al., 2016). Similarly, on a larger scale, the pathologist can know from the confidence intervals that for example at least 900 out of 1,000 images are myeloblasts. An example image of how the confidence intervals would be displayed is seen below.
One issue with current machine learning research for leukemia diagnosis has been the lack of prospective applications of algorithms in real-world scenarios (Salah et al., 2019). Researchers predominantly use small and homogenous samples to train models that provide point predictions without explanation. These are some of the issues we had in mind while designing our white blood cell identifier. We explore a methodology from Angelopoulos et al., to quantify the uncertainty of our models’ predictions and provide accurate confidence intervals so that pathologists can understand the probability of certain cells being present. This feature can be useful in real world scenarios in multiple ways. For example, given a cell to identify, a pathologist can know with 95% confidence that the cell presented is not a myeloblast, which could be a cell indicative of AML if found in high numbers (DiNardo et al., 2016). Similarly, on a larger scale, the pathologist can know from the confidence intervals that for example at least 900 out of 1,000 images are myeloblasts. An example image of how the confidence intervals would be displayed is seen below.
Figure 1: Example image of uncertainty quantification
Recent research on applying machine learning to leukemia has been focused on ALL, which is the most common form of leukemia diagnosed in children, a smaller number of studies focussed on AML, which is a common form in adults (Mayo Clinic, n.d.). Strategies included training models on genetic data, bone marrow cell morphology, and blood cell images (Nazari et al., 2020; Huang et al., 2020; Ahmed et al., 2019). In a paper titled, “Human-level recognition of blast cells in acute myeloid leukemia with convolutional neural networks,” researchers collect a dataset of single cell images of white blood cells, some from AML patients, and train a neural net for cell classification (Matek et al., 2019). They maintain a similar motivation to the goals of our project, stating: “Our approach holds the potential to be used as a classification aid for examining much larger numbers of cells in a smear than can usually be done by a human expert”. We worked to improve the methodology and applicability of this technology in a number of ways, including: building models that are more reliable, ensuring no data leakage, providing more information than just point predictions, and using more sound data augmentation techniques.